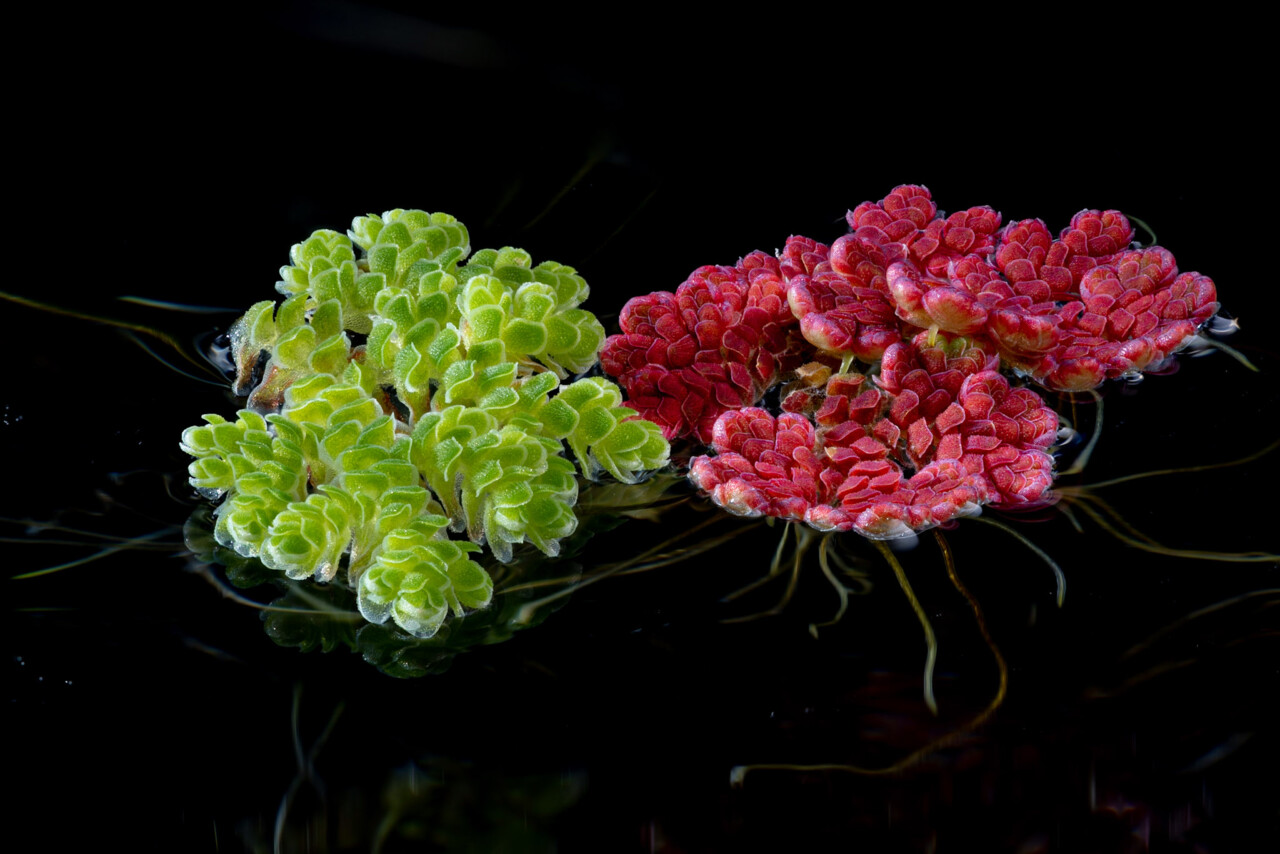
Hanneke Suijkerbuijk, chair
PhD candidate, Laboratory of Entomology, Wageningen University & Research
About my research
Insect herbivores such as caterpillars and aphids can cause great damage to plants in the field, with disastrous consequences for plant fitness in terms of seed yield and quality. Apart from direct damage to flowers and seeds, herbivores affect yield and seed quality indirectly by changing plant-pollinator interactions or by reducing outcrossing rates due to favouring self-pollination under stressful conditions. A major knowledge gap in both fundamental and applied aspects of seed yield and quality is how they are affected by these indirect interactions between herbivores and pollinators.
My work aims to gain more insight into how insect herbivory affects various pollination processes and how plants integrate defense and reproduction strategies. I perform field experiments to study the effects of herbivory on Brassica rapa seed set and the response of the pollinator community in terms of attraction and behaviour; greenhouse studies to look more closely at the effects of herbivory on male fitness and self-incompatibility; and laboratory studies for a more mechanistic understanding of plasticity in self-incompatibility.

Davar Abedini, secretary
PhD candidate, Plant Hormone Biology, University of Amsterdam
Project: Microbial recruitment by tomato and potato roots under nutrient deficiency
About my research
Under unfavorable conditions, plants exude a plethora of signaling molecules through complex series of biological mechanisms to recruit beneficial microbe(s). These beneficial microbes are involved in a range of processes from improving nutrient availability, providing growth hormones, and modulating the abiotic stresses response in plants to mitigating biotic stresses by inducing resistance and synthesizing antibiotics targeting pathogens. To facilitate this beneficial interaction, plants and their microbial partners have evolved a sophisticated chemical dialogue. Using small molecules, plants communicate with and modify the microbiome composition in their rhizosphere. My project aims to unravel potato/tomato-microbe chemical communications. For this, I am using multiomics approaches coupled with advanced analytical chemistry and molecular biology tools to identify the signalling molecules and their corresponding biosynthetic genes. The project will pave the ways toward reaching sustainable agriculture.

Thijs Bierman, council member
PhD candidate, Above-Belowground interactions, Institute of Biology, Leiden University
Project: Self Defense: Mimicking natural deterrent strategies in plants using adhesive spheres and volatiles
About my research
Arthropods such as insects and mites can be major pests in greenhouses. They eat our plants and can also transmit plant diseases. Chemical pesticides can be toxic for the environment, human health, or simply stop working due to pests becoming resistant. The Plant self-defense project that I take part in aims to develop a new crop protection method based on natural materials. By spraying tiny adhesive particles on the plant, we turn the whole plant into a sticky trap. The idea is to catch tiny arthropods, such as Western Flower thrips, our target pest. We work with crops such as chrysanthemum, tomato, and strawberry. Wageningen and Groningen universities produce the adhesive particles, while at Leiden we perform experiments to see if thrips damage is reduced by our new protection method and how the plant itself responds by showing growth damage or via changes in its metabolome. Together with Aeres Applied University we investigate effects on non-target insects, such as biocontrol agents of thrips and bumblebees. I also do research on volatiles that are attractive and repellent to thrips, which we would like to incorporate into the adhesive spheres to make them even more effective.

Robin Cowper, council member
PhD candidate, Plant-Microbe Interactions, Utrecht University
Project: Mycoat: Creating sustainable seed coatings
About my research
Seeds are the foundation of global food production. Their viability, germination and resistance to stress are key to feeding the world. Conventionally, plant breeders coat the surface of seeds with extra materials to improve their handling and vitality, a process known as coating . The ingredients in seed coatings include nutrients, herbicides, fungicides, and insecticides. Seed coating promotes the rapid and uniform germination of seeds, ensures their survival against abiotic and biotic stress factors, and ensures a high crop yield. Seed coating technology is emerging as an alternative tool to conventional farming because seed coating uses minor amounts of chemical inputs during its application (Rocha et al., 2019). Despite their effectiveness, synthetic pesticides and fertilisers can accumulate in plants, soil and water, causing toxicity to microbial populations. Thus, we need less toxic and more biodegradable coatings.
Plant beneficial microbes (PBMs) can be an alternative to the use of agrochemicals in plant production. PBMs help plants maintain or increase plant growth, unlock nutrients for the plant, and reduce crop loss caused by pathogens, insects and abiotic stress. For example, Trichoderma and Pseudomonas strains added to agricultural soils can antagonise pathogens and induce resistance in plants against bacterial, viral and fungal pathogens. In this project, the chair groups of PMI, Microbiology and NMR aim to develop sustainable seed coatings with plant-beneficial properties, based on fungal materials. We are looking at ways in which seed coating materials antagonize plant pathogens to protect emerging seeds from disease. Moreover, by using pathogen bioassays and digital plant phenotyping tools, we evaluate the contribution of our seed coatings to long-term plant health and resistance. We will also delve deeper into the modulation of the plant’s transcriptomic landscape and microbiome, when seeds interact with PBMs. This project can help us understand how fungal materials can be used in seed coating technology to support plant growth and resistance to pathogens.

Anouk Hendriks, council member
PhD candidate, Plant Breeding, Wageningen University & Research
Project: Characterization of a decreased resistance to downy mildew in nonhost Lactuca saligna.
About my research
One of the most harmful diseases on lettuce is downy mildew, caused by the oomycete pathogen Bremia lactucae. Bremia is an obligate biotroph and infects the leaves of its host, resulting in necrotic tissue, and therefore creating devastating yield losses in the fields. Current strategies for downy mildew control are mainly focused on fungicides and finding qualitative (race-sprecific and dominant) resistance (R) genes that establish a gene-for-gene interaction with avirulence (Avr) genes of specific Bremia strains. Over time, both methods of resistance prove to be ineffective due to the frequent formation of new B. lactucae strains. Alternative resistance mechanisms need to be identified and studied to provide a more durable resistance that is not easily overcome.
The wild lettuce species Lactuca saligna is a nonhost to B. lactucae and it is believed that its complete resistance is composed of such alternative resistance mechanisms. Due to the indicated race-nonspecific quantitative effects and the polygenic inheritance of this resistance, this type of layered resistance (QTLs) will not be easily overcome by the pathogen. In this project, a rare mild susceptible accession will be characterized. A novel source of quantitative B. lactucae resistance is, hereby, expected to be identified of which the genetics and role in the immunity network will be studied. Findings of this study are then expected to contribute to the understanding of the quantitative disease resistance of L. saligna, while the newly identified genes for resistance are expected to be used to improve the resistance in cultivated lettuce (L. sativa).

Sjors Huizinga, council member
PhD candidate, Plant Hormone Biology group, University of Amsterdam
Project: How do root-colonizing fungi sense plant roots?
About my research
Plants spend up to a third of their energy on the production of metabolites that are then exuded from the roots. These so-called root exudates act to shape the rhizosphere to the needs of the plant. For example, by modifying soil properties to be more favorable, but also by attracting and promoting the growth of certain microbes, leading to the establishment of a unique root microbiome. Some of these microbes are thought to be beneficial to the plant, whereas other can be detrimental. Root colonizing fungi are an important part of the root microbiome. Trichoderma species, for example, can benefit the plant by facilitating nutrient uptake and by attacking pathogenic fungi. Certain varieties of Fusarium species on the other hand, are pathogenic and can lead to devastating agricultural losses.
My work is focused on the signaling that occurs between lettuce and the root-colonizing fungi Trichoderma and Fusarium in the soil. I am trying to identify which molecules are present in lettuce root exudate, how the composition of this exudate changes during stress, and which molecules can elicit biological responses from the fungi. Simultaneously, I am trying to identify and characterize the receptors that the fungi use to perceive these signals.

Maxence Longuemare, council member
PhD candidate, Entomology, Wageningen University and Research
Project: Static but Plastic : Exploring plant defence strategies to deal with attack by multiple insect herbivores
About my research
In nature, plants are confronted with a plethora of insect herbivores which have been driving the evolution of plant defences for more than 400 million years. Plants have developed plastic defensive strategies to be resistant or tolerant against their diverse and, sometimes, unpredictable antagonist communities. While our understanding on plant-insect interactions has greatly improved over the past decade, most studies have focused on single or dual herbivore attack which do not capture the complexity of interactions occurring in natural ecosystems. Which growth-defence strategies plants may deploy to defend themselves under situation of attack by multiple herbivorous insects is largely unexplored. The aim of my PhD project is to investigate the plasticity of these defensive strategies within the Brassicaceae family, to understand the evolution of plant defences. Specifically, I explore how plants deal with sequential attack by multiple insects from different feeding guilds, how variation in time and space affects plant defence plasticity and how the different defence strategies vary between related plant species. For this, I am using transcriptomic and metabolic approaches, coupled with phenotypic measures on plant growth, reproduction, and insect performance.

Gyongyi Macias Honti, council member
PhD candidate at Plant Stress Resilience, Utrecht University & Plant Systems Physiology, Radboud University
Project: Mapping out site of action for ethylene-related root meristem hypoxia pre-adaptation
About my research
As climate change progresses, flooding events have become more frequent and intense. Flooding subjects plant roots to hypoxic (low oxygen) conditions, hindering gas exchange and leading to the accumulation of ethylene (volatile plant hormone), making this an important flood detection cue. Plant plasticity is especially important for plants in order to survive sub-optimal and stress conditions. Meristematic cells are key players in this because they are undifferentiated cells found at shoot and root tips with the potential to generate organs. The mechanisms behind meristem tolerance to stress conditions have yet remained underexplored. Previous experiments have shown that, in Arabidopsis, an ethylene pre-treatment following subsequent hypoxic stress can enhance root survival. While it is clear that the activating mechanism caused by ethylene accumulation boosts the regrowth capacity of roots after hypoxia, it is yet unknown how meristem protection is conferred. Is ethylene mediated by specific cell layers in the root? Do specific cell layers communicate signals in order to coordinate meristem protection?
My project focuses on elucidating the role of ethylene in conferring meristematic protection and to understand where exactly this is carried out in the root. For this, I use cell-type specific lines that have a disruption in ethylene signaling, and through imitating flooding conditions, study the survival effect on root tips. This is coupled with other reporter lines in order to be able to further understand what other players are involved in this meristematic protection during flooding conditions.

Sergio Martín-Ramírez , council member
PhD candidate, Laboratory of Biochemistry, Wageningen University & Research
Project: Redox-dependent extracellular interaction networks of Cysteine-Rich- and Leucine-Rich Repeat- Receptor Kinases.
About my research
Reactive Oxygen Species (ROS) are produced in intra- and extra-cellular compartments as response for any biotic and abiotic stresses. ROS also act as signal molecules in development and defence, however, the mechanisms and receptors by which extracellular ROS signals are perceived and integrated are still unclear. We propose that Cysteine-Rich Receptor-Like Kinases (CRKs), a family of Receptor Kinases (RKs) in Arabidopsis, can serve as extracellular ROS sensors. We propose that ROS modulate interactions between CRKs and Leucine-Rich Repeat Receptor Kinases (LRR-RKs) reflecting the redox state of the apoplast. We aim to elucidate how RKs signalling networks are modulated by ROS, and how this orchestrates responses to stresses by creating proteome-wide networks of interactions among extracellular domains (ECDs) of RKs in vitro.
The first part of the project will consist of building a Redox-dependent Interaction Network between the ECDs of LRR-RKs and CRKs of Arabidopsis using a large-scale interactome screen in different redox conditions (Smakowska-Luzan et al., 2018). The second part will assign a biological relevance to the redox-dependent CRKs interactions. Relevant candidates for redox-modulated interactions will be selected for further in-depth functional characterization using biochemical, molecular and cell biology approaches.

Sanne Matton, council member
PhD candidate, Plant-Environment Signaling, Utrecht University & Wageningen University & Research
About my research
Plants are able to perceive the quality and quantity of the light in their environment using several different light receptors, which enables them to respond adequately to their light environment and optimize light harvesting for photosynthesis. Via their phytochrome photoreceptors plants are able to detect shade. As a consequence, when a shade avoiding plant like Arabidopsis thaliana perceives shade its energy investment shifts away from processes like growth of the leaf lamina, root system and fruits towards an increased elongation of the stem and upwards growth of leaves. The shade avoidance response, which is also present in many crops, thus affects yield of edible and usable parts of said crops.
The shade avoidance response has been researched thoroughly in seedlings, and already some work has been done on adult plants. In my project I focus on adult plants, further elucidating growth responses upon spatial and temporal fluctuations in light quality and quantity in Arabidopsis thaliana and Solanum lycopersicum. For example monitoring leaf growth responses when only a small part of the leaf is shaded, but also investigating the effect of fluctuations in the light quality or quantity over time. The aim of my project will be to elucidate the molecular pathways underlying local light dependent growth responses and the effect spatial and temporal fluctuations in light quality and quantity have on leaf growth and development.

Gabriele Panicucci, council member
PhD candidate, Plant-Environment Signaling, Utrecht University
About my research
The abundance of molecular oxygen in our atmosphere allows plants to efficiently produce energy through mitochondrial respiration. Plants experiencing oxygen-limiting conditions, such as flooding and waterlogging, have to cope with these sudden energetic constrains by relying on fermentative metabolism and species-specific morphological adaptations. Therefore, scarcity of oxygen has traditionally been studied in the context of abiotic stress.
Surprisingly, meristematic tissues in plants were found to experience a condition of chronically low oxygen levels regardless of external oxygen availability. Specifically, stem cells hosted within the shoot apical meristem were shown to thrive in a state of constant hypoxia. In my research I mostly investigate the link between oxygen distribution, low-oxygen signaling and meristematic activity.

Alan Pauls, council member
PhD candidate, Genetics, Wageningen University & Research
Project: LettuceKnow Project 2.2 “Genetics of abiotic stress resilience in lettuce”
About my research
It is well known that environmental factors such as light intensity, salinity levels and nutrient availability play an important role in plant growth and yield, yet the molecular mechanisms and genetic architecture that underline the responses of plants to said conditions are still largely unknown. With climate change causing erratic weather patterns, arable land with ideal environmental conditions is becoming an increasingly scarce resource. This necessitates the need to identify stress resilience loci and develop stress resistant crops. An important, yet underutilized avenue that can be used for identification of novel stress resilience loci is by mining the extended germplasm of crops thus exploring and exploiting its existing natural genetic variation. This approach has been used extensively in Arabidopsis thaliana but in the case of lettuce the use has been limited to the search for immunity related traits. With the advent of high throughput phenotyping technologies that can phenotype a large number of plants continuously across multiple days, phenotyping complex traits linked to abiotic stress can be more robustly captured. This, combined with powerful integrative bioinformatics and machine learning could help describe the genetic background of abiotic stress resilience in lettuce.
In a nutshell, my project a part of the larger LettuceKnow project aims “To identify and exploit natural variation in the LK500 population to improve lettuce resilience to abiotic stress conditions specifically fluctuating light and tipburn inducing growth conditions while reducing the trade-off towards plant growth”.

Sanne Put, council member
PhD candidate, Plant Breeding, Wageningen University & Research
Project: Breeding for improved yield and attachment of the sugar kelp Saccharina latissima
About my research
We currently need almost half of the land surface to produce plant and animal based food and this percentage has to increase to ensure food security in the future. The world population continues to grow and simultaneously agricultural yield is expected to decrease due to soil degradation, exhaustion of natural resources and extreme weather events. The sea provides opportunity to produce seaweed for food without relying on irrigation with fresh water, high quality arable land, and in some cases fertilisers. However, seaweed farming in the North Sea is not yet economically feasible. The challenge to make the seaweed sector profitable is two-fold – unpredictable, low yields and high production costs. Limited breeding efforts for Saccharina latissima, the sugar kelp species native to the North Sea, hampered improvement of yield and yield stability. There is currently little understanding of the genetics determining complex yield traits and there are limited tools available for high-throughput screening of yield of S. latissima. By high-throughput screening the natural variation that exists we can unravel yield related physiological processes that evolved naturally. Besides low yields, production costs associated with the current cultivation method predominantly used in Asia are high. To reduce production costs, the European seaweed sector wants to shift to a so-called direct seeding method, however farmers have observed high losses of up to 99% of the seeding material. To reduces these losses, attachment of seaweed to the ropes should be enhanced. By acquiring knowledge of the genetics underlying the complex traits yield and attachment, breeders will be able to provide farmers with more robust starting material. During my PhD, I will develop a high-throughput phenotyping method for yield and perform a genome-wide association study (GWAS) for yield and attachment with over 200 different S. latissima genotypes from 34 locations.

Sophie Vijverberg, council member
PhD candidate, Plant-Microbe Interactions, Utrecht University
Project: The identification of candidate genes in the interaction between Arabidopsis and plant-growth promoting rhizobacteria.
Beneficial microbes in the microbiome of plants can serve as their allies, by stimulating plant growth and enhancing defense against pathogens. These beneficial microbes are collectively referred to as plant-growth promoting rhizobacteria (PGPR). The PGPR Pseudomonas simiae WCS417 can colonize Arabidopsis thaliana roots, resulting in morphological changes in both its roots and shoots. These changes include an increase in shoot fresh weight, alongside an increase in primary root length and the number of lateral roots. It has been established that not all Arabidopsis accessions respond equally to WCS417; the morphological changes in some accessions are much more pronounced than in others. This indicates natural variation in the ability of Arabidopsis to benefit from an interaction with WCS417.
The aim of my project is to improve our understanding of what Arabidopsis genes are important in its response to WCS417. We expect to find candidate genes by conducting a Genome-wide association study (GWAS) on a large collection of natural Arabidopsis accessions. I will use the high-throughput imaging facilities from NPEC to generate the large amount of phenotypic data required for the GWAS. My research may contribute to the development of future crops optimized in benefiting from the positive effect from PGPR’s.

Elmar van der Wijk, council member
PhD candidate, Plant Systems Physiology, Radboud University & Plant-Environment Signaling, Utrecht University
Project: Unravelling the cell-type-specific mechanisms of ethylene priming for meristem protection against hypoxia
About my research
During flooding, reduced gas exchange subjects plants to reduced oxygen levels (hypoxia). This also causes a build-up of the volatile plant hormone ethylene, which acts as an early warning signal. Plants treated with ethylene show increased hypoxia tolerance. Central in this pathway is the ethylene-induced gene phytoglobin 1 (PGB1). PGB1 removes nitric oxide (NO) from the submerged plant root, and thereby stabilises the group-VII ethylene response factor (ERF-VII) transcription factors that in turn activate genes that protect the meristem against subsequent hypoxia. Despite the established importance of priming against hypoxia, little is known about the mechanisms of this meristem protection by ethylene and phytoglobin.
During my PhD, in collaboration with colleagues at Utrecht University, we aim to elucidate the regulation of PGB1, and the cell-type-specific response to ethylene and hypoxia. For this, we will study the importance of each cell type in this process and investigate the spatiotemporal regulation of the central genes. During my PhD, I use an integrated approach that incorporates both experimental data and bioinformatics analyses. Finally, this project paves the way for the creation of more flooding stress-resilient crops.